Chapter 1 End Problems
Problem 3
Aa X Aa produces 6 offspring, so the number of trials is 6.
Expect aa 25% of the time
and expect not aa 75% of the time.
1 | probability.of.two <-(factorial(6)/(factorial(4)*factorial(2)))*(0.75^4)*(0.25^2) |
Let’s try a brute force approach. Populate a vector of length 1000 with 250 AA, 500 Aa and 250 aa. Sample with replacement 1000 times 6 geneotype. Calculate the fraction with aa present 2 or fewer times.
1 | population <- c( rep("AA", 250), rep("Aa", 500), rep("aa", 250)) |
## Problem 4
Jack, queen, king are the face cards so Jack is likely 1/3 of the time a face card is drawn.
There are 4 Jacks so Jack of Hearts is 1 of 4.
The probability that a randomly drawn face card (of which there are 12) is the jack of hearts is 1 of 12.
Figure out potentially relevant probabilities then answer the question.
P{FaceCard}: probability of a Face card = 12/52 = 3/13
P{Jack}: probability of a Jack = 4/52 = 1/13.
P{Hearts}: probability of a hearts = 13/52 = 1/4.
P{Jack | FaceCard}: probability of a Jack given it’s a face card = 4/12 = 1/3.
P{Jack | Hearts}: probability of a Jack given it’s a heart = 1/13.
P{Heart | Jack}: probability of a heart given a Jack = 1/4
P{FaceCard | Jack}: probability of a face card given it’s a Jack = 1
What is the probability that a randomly drawn face card is the jack of hearts?
P{JackHearts | FaceCard} = P{ Heart | Jack } X P{Jack | FaceCard}
= 1/4 x 1/3 = 1/12
Perform a trivial Bayesian analysis
Bayes Rule: P{A | B} = P{B | A}*P{A}/P{B}
What is the probability of a Jack given a face card?
There are 12 Face cards and 4 are Jacks so 4/12 or 1/3
Rewrite the Bayesian rule as: P{Jack|FaceCard} = P{FaceCard|Jack}*P{Jack}/P{FaceCard}
P{Jack|FaceCard} = (1*1/13)/3/13 = 1/3
## Problem 6
1 | observed <- c( 60, 40) |
r = 40/100 = 0.4
standard error = sqrt(( 0.4*( 1-0.4 ))/100 = 0.049
## Problem 12
This R solution requires the package “seqinr”
1 | template <- "tactgtgaaagatttccgact" |
For a plotting example using seqinr, see
## Problem 13
1 | K <- 100000 |
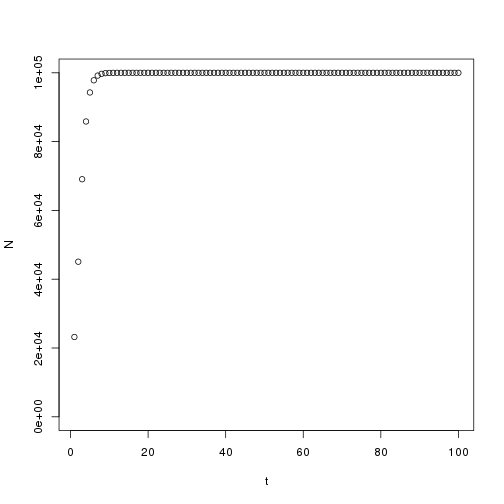
1 | #Let's change the rate to 0.1 |

1 | #It takes longer to reach the carrying capacity |

1 | #The population crashes to 0 |

1 | #Population drops to the carrying capacity |
## Problem 14
1 | observed <- c(88, 37) |
## Problem 15
1 | observed <- c(269, 98, 86, 88, 30, 34, 27, 7) |